A rational and systematic approach to PROTACs development: identification of CBP/EP300 degraders
PROteolysis TArgeting Chimeras (PROTACs) are bifunctional molecules that simultaneously bind an E3 ligase and a protein of interest, inducing degradation of the latter via the ubiquitin-proteasome system. Despite their outstanding potential, the rational design of PROTACs remains challenging as several variables need to be tuned in parallel to obtain active degraders, and their large size and novel structure activity relationship (SAR) means that much of the knowledge surrounding the design of small molecule drugs cannot be applied. To tackle this problem, we systematically explored PROTAC SAR, leading to the rational design of degraders targeting CREB-binding protein (CBP) and E1A-associated protein (EP300) - two homologuous regulatory proteins crucial for enhancer mediated transcription and implicated in a wide range of human diseases. We established a novel synthetic route to improve the chemical accesibility of the CBP/EP300 ligand, and several cell-based assays including the measurment of cellular ternary complex formation for the first time for Cereblon (CRBN) and CBP. Through this we demonstrate that engagement of CRBN, rather than VHL or IAP, and a specific linker between the E3 and CBP binders are essential for PROTAC-mediated CBP degradation, and by doing so, we broaden the tools available to modulate CBP and EP300. Lessons learnt from this campaign, particularly the importance of cell-based assays beyond degradation to understand reasons for PROTAC inactivity, are widely applicable to other challenging targets and can assist in guiding the development of degraders whilst minimizing compound libraries.
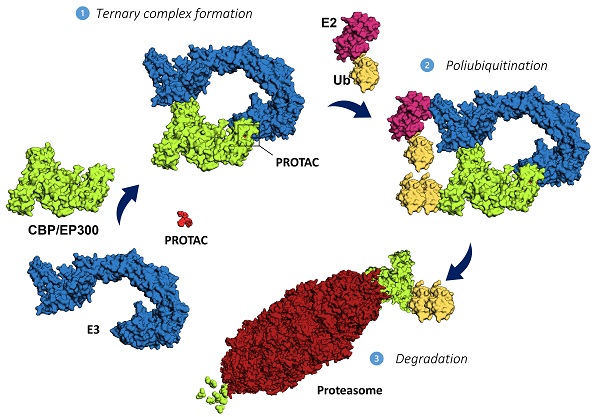
[2] Ashton C. Lai, Craig M. Crews, Nature Reviews Drug Discovery, 2017, 16, 101-114
[3] Taku Watanabe et al., Scientific Reports, 2017, 7, 46380.